Similarity measures¶
This section describes the various similiarity measures implemented in mermaid.
Similarity measure factory¶
This is a factory which generates all the supported similarity measures.
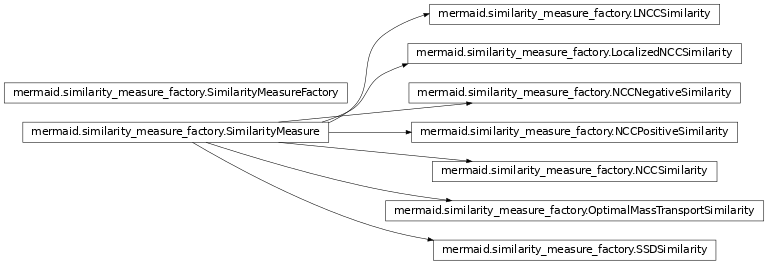
Similarity measures for the registration methods and factory to create similarity measures.
-
class
mermaid.similarity_measure_factory.SimilarityMeasure(spacing, params)[source]¶ Abstract base class for a similarity measure.
-
spacing= None¶ pixel/voxel spacing
-
volumeElement= None¶ volume element
-
dim= None¶ image dimension
-
params= None¶ external parameters
-
sigma= None¶ 1/sigma^2 is a balancing constant
-
compute_similarity_multiNC(I0, I1, I0Source=None, phi=None)[source]¶ Compute the multi-image multi-channel image similarity between two images of format BxCxXxYzZ
Parameters: - I0 – first image (the warped source image)
- I1 – second image (target image)
- I0Source – source image (will typically not be used)
- phi – map in the target image to warp the source image (will typically not be used)
Returns: returns similarity measure
-
compute_similarity_multiC(I0, I1, I0Source=None, phi=None)[source]¶ Compute the multi-channel image similarity between two images of format CxXxYzZ
Parameters: - I0 – first image (the warped source image)
- I1 – second image (target image)
- I0Source – source image (will typically not be used)
- phi – map in the target image to warp the source image (will typically not be used)
Returns: returns similarity measure
-
compute_similarity(I0, I1, I0Source=None, phi=None)[source]¶ Abstract method to compute the single-channel image similarity between two images of format XxYzZ. This is the only method that should be overwritten by a specific implemented similarity measure. The multi-channel variants then come for free.
For proper implementation it is important that the similarity measure is muliplied by 1./(self.sigma ** 2) and also by self.volumeElement if it is a volume integral (and not a correlation measure for example)
Parameters: - I0 – first image (the warped source image)
- I1 – second image (target image)
- I0Source – source image (will typically not be used)
- phi – map in the target image to warp the source image (will typically not be used)
Returns: returns similarity measure
-
-
class
mermaid.similarity_measure_factory.SSDSimilarity(spacing, params)[source]¶ Sum of squared differences (SSD) similarity measure.
\(1/sigma^2||I_0-I_1||^2\)
-
class
mermaid.similarity_measure_factory.OptimalMassTransportSimilarity(spacing, params, sinkhorn_iterations=200, std_sinkhorn=0.08)[source]¶ - Constructor
param spacing: spacing of the grid (as in parameters structure) param std_dev: similarity measure parameter param sinkhorn_iterations: number of iterations in sinkhorn algorithm param std_sinkhorn: standard deviation of the entropic regularization (not too small to avoid nan)
-
spline_order= None¶ order of spline for interpolation (if needed)
-
class
mermaid.similarity_measure_factory.NCCSimilarity(spacing, params)[source]¶ Computes a normalized-cross correlation based similarity measure between two images. \(sim = (1-ncc^2)/(\sigma^2)\)
-
class
mermaid.similarity_measure_factory.NCCPositiveSimilarity(spacing, params)[source]¶ Computes a normalized-cross correlation based similarity measure between two images. Only allows positive correlations. \(sim = (1-ncc)/(\sigma^2)\)
-
class
mermaid.similarity_measure_factory.NCCNegativeSimilarity(spacing, params)[source]¶ Computes a normalized-cross correlation based similarity measure between two images. Only allows negative correlations. \(sim = (ncc)/(\sigma^2)\)
-
class
mermaid.similarity_measure_factory.LNCCSimilarity(spacing, params)[source]¶ This is an generalized LNCC; we implement multi-scale (means resolution) multi kernel (means size of neighborhood) LNCC.
Param: resol_bound : type list, resol_bound[0]> resol_bound[1] >… resol_bound[end] Param: kernel_size_ratio: type list, the ratio of the current input size Param: kernel_weight_ratio: type list, the weight ratio of each kernel size, should sum to 1 Param: stride: type_list, the stride between each pixel that would compute its lncc Param: dilation: type_list Settings in json:
"similarity_measure": { "develop_mod_on": false, "sigma": 0.5, "type": "lncc", "lncc":{ "resol_bound":[-1], "kernel_size_ratio":[[0.25]], "kernel_weight_ratio":[[1.0]], "stride":[0.25,0.25,0.25], "dilation":[1] }
For multi-scale multi kernel, e.g.,:
"resol_bound":[64,32], "kernel_size_ratio":[[0.0625,0.125, 0.25], [0.25,0.5], [0.5]], "kernel_weight_ratio":[[0.1,0.3,0.6],[0.3,0.7],[1.0]], "stride":[0.25,0.25,0.25], "dilation":[1,2,2] #[2,1,1]
or for single-scale single kernel, e.g.,:
"resol_bound":[-1], "kernel_size_ratio":[[0.25]], "kernel_weight_ratio":[[1.0]], "stride":[0.25], "dilation":[1]
Multi-scale is controlled by “resol_bound”, e.g resol_bound = [128, 64], it means if input size>128, then it would compute multi-kernel lncc designed for large image size, if 64<input_size<128, then it would compute multi-kernel lncc desiged for mid-size image, otherwise, it would compute the multi-kernel lncc designed for small image. Attention! we call it multi-scale just because it is designed for multi-scale registration or segmentation problem. ONLY ONE scale would be activated during computing the similarity, which depends on the current input size.
At each scale, corresponding multi-kernel lncc is implemented, here multi-kernel means lncc with different window sizes Loss = w1*lncc_win1 + w2*lncc_win2 … + wn*lncc_winn, where /sum(wi) =1 for example. when (image size) S>128, three windows sizes can be used, namely S/16, S/8, S/4. for easy notation, we use img_ratio to refer window size, the example here use the parameter [1./16,1./8,1.4]
In implementation, we compute lncc by calling convolution function, so in this case, the [S/16, S/8, S/4] refers to the kernel size of convolution function. Intuitively, we would have another two parameters, stride and dilation. For each window size (W), we recommend using W/4 as stride. In extreme case the stride can be 1, but can large increase computation. The dilation expand the reception field, set dilation as 2 would physically twice the window size.
-
class
mermaid.similarity_measure_factory.LocalizedNCCSimilarity(spacing, params)[source]¶ Computes a normalized-cross correlation based similarity measure between two images. \(sim = (1-ncc^2)/(\sigma^2)\)
-
gaussian_std= None¶ half the side length of the cube over which lNCC is computed
-
-
class
mermaid.similarity_measure_factory.SimilarityMeasureFactory(spacing)[source]¶ Factory to quickly generate similarity measures that can then be used by the different registration algorithms.
-
spacing= None¶ image spacing
-
dim= None¶ dimension of image
-
similarity_measure_default_type= None¶ default image similarity measure
-
simMeasures= None¶ currently implemented similiarity measures
-
add_similarity_measure(simName, simClass)[source]¶ Adds a new custom similarity measure
Parameters: - simName – desired name of the similarity measure
- simClass – similiarity measure class (whcih can be instantiated)
-
set_similarity_measure_default_type_to_omt()[source]¶ Set the default similarity measure to OMT (optimal mass transport)
-
set_similarity_measure_default_type_to_ncc_positive()[source]¶ Set the default similarity measure to positive NCC (i.e., only positive correlations allowed)
-
set_similarity_measure_default_type_to_ncc_negative()[source]¶ Set the default similarity measure to positive NCC (i.e., only negative correlations allowed)
-
Similarity helper OMT¶
These are some helper functions for the optimal mass transport similarity measure.
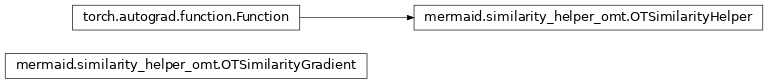
Similarity measures for the registration methods and factory to create similarity measures.
-
class
mermaid.similarity_helper_omt.OTSimilarityHelper[source]¶ Implements the pytorch function of optimal mass transport.
-
static
forward(ctx, phi, I0, I1, multiplier0, multiplier1, spacing, nr_iterations_sinkhorn, std_sink)[source]¶ Performs the operation.
This function is to be overridden by all subclasses.
It must accept a context ctx as the first argument, followed by any number of arguments (tensors or other types).
The context can be used to store tensors that can be then retrieved during the backward pass.
-
static
backward(ctx, grad_output)[source]¶ Defines a formula for differentiating the operation.
This function is to be overridden by all subclasses.
It must accept a context
ctxas the first argument, followed by as many outputs didforward()return, and it should return as many tensors, as there were inputs toforward(). Each argument is the gradient w.r.t the given output, and each returned value should be the gradient w.r.t. the corresponding input.The context can be used to retrieve tensors saved during the forward pass. It also has an attribute
ctx.needs_input_gradas a tuple of booleans representing whether each input needs gradient. E.g.,backward()will havectx.needs_input_grad[0] = Trueif the first input toforward()needs gradient computated w.r.t. the output.
-
static
-
class
mermaid.similarity_helper_omt.OTSimilarityGradient(spacing, shape, sinkhorn_iterations=300, std_dev=0.07)[source]¶ Computes a regularized optimal transport distance between two densities.
Formally: \(sim = W^2/(\sigma^2)\)
-
build_kernel_matrix(length, std)[source]¶ Computation of the gaussian kernel.
Parameters: - length – length of the vector
- std – standard deviation of the gaussian kernel
Returns: \(\exp(-|x_i - x_j|^2/\sigma^2)\)
-
build_kernel_matrix_gradient(length, std)[source]¶ Computation of the gaussian first derivative kernel multiplied by \(1/2\sigma^2\)
Parameters: - length – length of the vector
- std – standard deviation of the gaussian kernel
Returns: \((x_i - x_j) \exp(-|x_i - x_j|^2/\sigma^2)\)
-
kernel_multiplication(multiplier)[source]¶ Computes the multiplication of a d-dimensional vector (d = 1,2 or 3) with the gaussian kernel K :param multiplier: the vector :param choice_kernel: the choice function that outputs the index in the kernel list. :return: K*multiplier
-
kernel_multiplication_gradient_helper(multiplier, choice_kernel)[source]¶ - Computes the multiplication of a d-dimensional vector (d = 1,2 or 3) with the
- (derivative along a given axis) gaussian kernel and given by the choice_kernel function (give the axis).
Parameters: - multiplier – the vector
- choice_kernel – the choice function that outputs the index in the kernel list.
Returns: K*multiplier
-
set_choice_kernel_gibbs(i, offset)[source]¶ Set the choice of the kernels for the computation of the gradient.
Parameters: - i – the (python) index of the dimension
- offset – the dimension
Returns: the function for choosing the kernel
-
compute_similarity(I0, I1)[source]¶ Computes the OT-based similarity measure between two densities.
Parameters: - I0 – first density
- I1 – second density
Returns: W^2/sigma^2
-
compute_gradient(I0, I1, multiplier0, multiplier1)[source]¶ Compute the gradient of the similarity with respect to the grid points
Parameters: - I0 – first density
- I1 – second density
- multiplier0 – Lagrange multiplier for the first marginal
- multiplier1 – Lagrange multiplier for the second marginal
Returns: Gradient wrt the grid
-